This website was created as an assignment for Genetics 564 at the University of Wisconsin-Madison
What is a Protein Domain and Motif?
A protein domain is a section of amino acid sequence that contains an important structural or functional unit of the overall protein. Domains are often conserved throughout evolution due to their importance to biological function of the protein and the organism itself [1]. Understanding the function of a domain can help scientists better understand disease and mutations related to the individual protein.
Motifs are usually smaller sections of the amino acid sequence that are responsible for the structure and folds of the protein or can be locations of activation or binding. These motifs are critical for other functions of the protein as well. Motifs are usually found within a protein domain [11].
Domains of Ryanodine Receptor 2
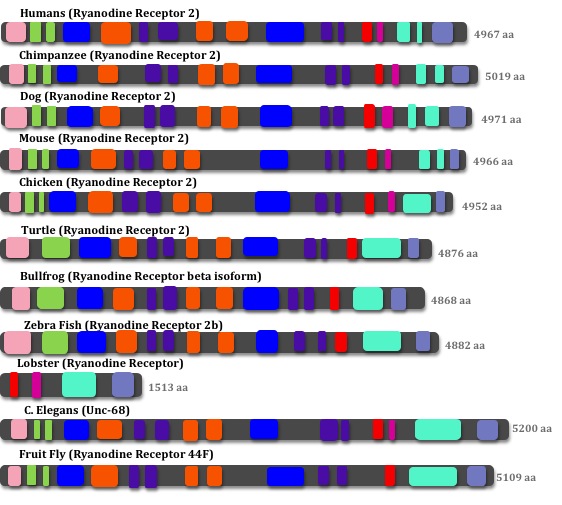
Since the Ryanodine Receptor 2 protein is so large and interacts with many other different proteins within cardiac tissue, it has many different domains. The domains outlined below were found using a combination of InterPro, SMART, Pfam, Uniprot, and HomoloGene.
MIR
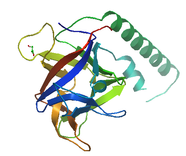
The MIR domain is named after the three proteins it is present in; Mannosyltransferases, Inositol 1,4,5-trisphosphate receptor, and the Ryanodine Receptor. MIR is localized in the membrane and is predicted to have ligand transferase function, but its function is still very unclear. The structure of this domain includes a closed beta-barrel structure with a hairpin triplet [2].
RYDR_ITPR
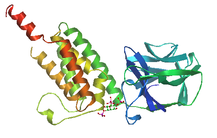
The RYDR_ITPR domain, commonly shorted to RIH domain, is found in calcium ion channel proteins. This domain is localized to the membrane and is involved in transmembrane calcium ion transport [3].
SPRY
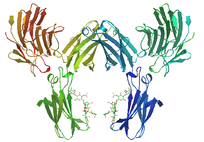
The SPRY domains are located in Spla and Ryanodine Receptors. Their function is still unknown[4].
RYR
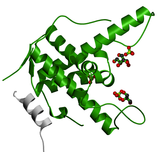
The RYR domain is found in Ryanodine Receptors. It is found in four different locations throughout the protein but the function of this domain is unknown [5].
RIH_Assoc
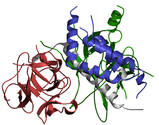
This domain is found in the protein family homotetrameric ligand-gated intracellular calcium ion channels, but has no known function[6].
EFh
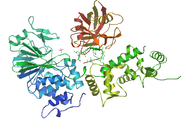
The EFh domain is involved in calcium-ion binding. This domain usually contains two alpha helices and a twelve residue loop connecting the two. When calcium binds to this domain, it changes conformation of the protein[7].
RR_TM4-6
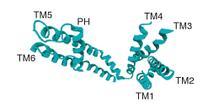
This ryanodine receptor TM4-6 domain is involved cellular calcium ion homeostasis. It uses a ryanodine induced calcium-ion release mechanism to pump calcium-ions out of the sacroplasmic reticulum[8].
Ins_143_p3
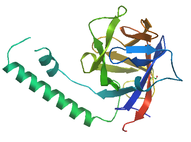
This region is usually located near the N-terminis of Ryanodine Receptors[9].
ion_trans
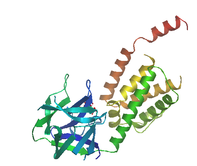
This domain is found in many different ion channels proteins including sodium, potassium, and calcium channels. It contains six helices and are involved in ion selectivity for the channel protein[10].
RYR2 Domains of Homologous Species
Motif
PROSITE identified 11 different motifs in the Ryanodine Receptor 2 protein. Many of these motifs occur in multiple regions. The following motif's were identified in RYR2:
N-myristoylation
Casein kinase II phosphorylation motif
Protein kinase C phosphorylation
N-glycosylation site
Tyrosine kinase phosphorylation site
Leucine zipper
Amidation site
cAMP- and cGMP-dependent protein kinase phosphorylation site
Cell attachment motif.
Click on the motifs above to learn more about their functions.
Analysis
Determining a protein's domains and motifs can help us better understand the overall function of a protein. This is particularly useful in genetic disorders for the understanding of the disorder as a whole. If a disease causing mutation is found within a domain or a motif, it can help us understand what function of the protein is being disrupted. Unfortunately, several of the domains of the RYR2 proteins have unknown functions. Further investigation of these unknown domains could provide insight that may be useful for finding a cure.
Determining a protein's domains and motifs can help us better understand the overall function of a protein. This is particularly useful in genetic disorders for the understanding of the disorder as a whole. If a disease causing mutation is found within a domain or a motif, it can help us understand what function of the protein is being disrupted. Unfortunately, several of the domains of the RYR2 proteins have unknown functions. Further investigation of these unknown domains could provide insight that may be useful for finding a cure.
References:
[1] EMBL-EBI. What are protein domains? retrieved Apr. 10, 2014. http://www.ebi.ac.uk/training/online/cours/introduction-
protein-classification-ebi/protein-classification/what-are-protein-domains
[2] MIR Domain-http://www.ebi.ac.uk/interpro/entry/IPR016093
[3] RIH Domain- http://pfam.sanger.ac.uk/family/PF01365#interproContent
[4] SPRY Domain- http://www.ebi.ac.uk/interpro/entry/IPR003877?q=SPRY
[5] RYR Domain- http://www.ebi.ac.uk/interpro/entry/IPR003032?q=RYR
[6] RIH_assoc Domain- http://www.ebi.ac.uk/interpro/entry/IPR013662?q=RIH_Assoc
[7] EF-Hand Domain-http://www.ebi.ac.uk/interpro/entry/IPR002048?q=EFh
[8] Ryanodine Receptor Trans Membrane Domain- http://www.ebi.ac.uk/interpro/entry/IPR009460
[9] Ins_143_p3 Domain- http://www.ebi.ac.uk/interpro/entry/IPR014821
[10] Ion_Trans Domain- http://www.ebi.ac.uk/interpro/entry/IPR005821?q=Ion_trans
[11] D'haeseleer, Patrik. (2006). What are DNA sequence motifs?. Nature Biotechnology.24. 423-425.
doi:10.1038/nbt0406-423 http://www.nature.com/nbt/journal/v24/n4/full/nbt0406-423.html
Prosite:http://prosite.expasy.org/
[1] EMBL-EBI. What are protein domains? retrieved Apr. 10, 2014. http://www.ebi.ac.uk/training/online/cours/introduction-
protein-classification-ebi/protein-classification/what-are-protein-domains
[2] MIR Domain-http://www.ebi.ac.uk/interpro/entry/IPR016093
[3] RIH Domain- http://pfam.sanger.ac.uk/family/PF01365#interproContent
[4] SPRY Domain- http://www.ebi.ac.uk/interpro/entry/IPR003877?q=SPRY
[5] RYR Domain- http://www.ebi.ac.uk/interpro/entry/IPR003032?q=RYR
[6] RIH_assoc Domain- http://www.ebi.ac.uk/interpro/entry/IPR013662?q=RIH_Assoc
[7] EF-Hand Domain-http://www.ebi.ac.uk/interpro/entry/IPR002048?q=EFh
[8] Ryanodine Receptor Trans Membrane Domain- http://www.ebi.ac.uk/interpro/entry/IPR009460
[9] Ins_143_p3 Domain- http://www.ebi.ac.uk/interpro/entry/IPR014821
[10] Ion_Trans Domain- http://www.ebi.ac.uk/interpro/entry/IPR005821?q=Ion_trans
[11] D'haeseleer, Patrik. (2006). What are DNA sequence motifs?. Nature Biotechnology.24. 423-425.
doi:10.1038/nbt0406-423 http://www.nature.com/nbt/journal/v24/n4/full/nbt0406-423.html
Prosite:http://prosite.expasy.org/
Site created by: Mercede Davis
Email contact: [email protected]
Date last updated: 5/14/14
Genetics 564, University of Wisconsin-Madison
Email contact: [email protected]
Date last updated: 5/14/14
Genetics 564, University of Wisconsin-Madison