This website was created as an assignment for Genetics 564 at the University of Wisconsin-Madison
What is Phylogeny?
Phylogeny is the study of the evolutionary paths of modern day species and their relationships based on a common ancestry. Scientists are able to use this information such as morphology, genetics, and the molecular biology of species to determine where they and their common relatives diverged from each other in the evolutionary process[1,2].
Phylogenetic Trees
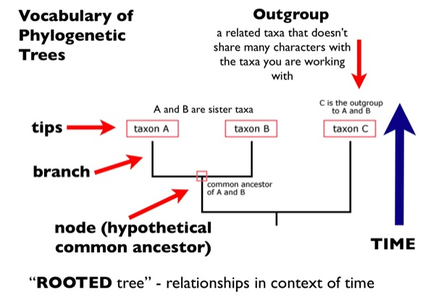 Figure 1: How to construct a phylogenetic tree [2]
Figure 1: How to construct a phylogenetic tree [2]
Scientists are able to use this information to construct a visual representation of this evolutionary process called a phylogenetic tree. Just like real trees, phylogenetic trees start from a common source called a root. From the root, there are two branches. As time goes on, the original two branches at the base of the tree continue to split and diverge creating a node at each division. As the tree's branches continue to split over and over as time goes on, more species appear until we search present day at the tips of the tree where all modern day organisms are represented. The species' lineage can be traced back down the tree and one can see where species diverged from each other by locating the node that separates them. These nodes represent a common ancestor shared between two or more species. The less nodes between organisms, the closer they are related[2]. For example, it is well known that chimpanzees and humans are closely related, though it is a common misconception that humans evolved from chimpanzees. On a phylogenetic tree, chimpanzees and humans are very close and very few nodes separate the two species. At the shared node, represents an organism very far back in history that both chimpanzees and humans evolved from. This ancestor would have had characteristics that were shared by both humans and chimpanzees. This concept will be seen below when we construct phylogenetic trees based only on the amino acid sequence of the RYR2 protein and its homologues.
Phylogeny of the RYR2 protein
Creating phylogenetic trees can be very easy using amino acid sequence. Sequences are inputted into an alignment program, in this case Clustal Omega was used again, and uses an algorithm to predict the evolutionary relationships between the organisms inputted and constructs the tree.
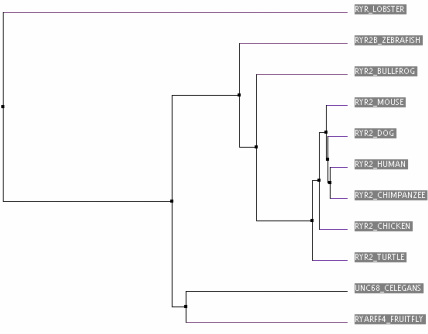
Trees can be constructed by using percent identity or by BLOSUM62's matrix. Percent identity trees uses the protein sequences and how similar they are to construct the tree.BLOSUM62 looks at each amino acid and determines how likely that amino acid is to mutate for a different amino acid for constructing it's trees[3].
Phylogeny of the RYR2 protein
Creating phylogenetic trees can be very easy using amino acid sequence. Sequences are inputted into an alignment program, in this case Clustal Omega was used again, and uses an algorithm to predict the evolutionary relationships between the organisms inputted and constructs the tree.
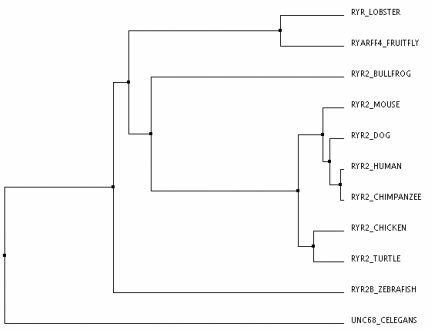
Trees can be constructed by using percent identity or by BLOSUM62's matrix. Percent identity trees uses the protein sequences and how similar they are to construct the tree.BLOSUM62 looks at each amino acid and determines how likely that amino acid is to mutate for a different amino acid for constructing it's trees[3].
Analysis
As you can see above by looking at the two trees, there is a lot of variability in phylogenetics. Both trees were constructed using a different process and produced two very different outcomes. On the tree created by BLOSUM62, the lobster is seen as the outgroup but on the tree created by percent identity, C. elegans is the outgroup. This discrepancy can probably be explained by the fact that the lobster's RYR protein is only about 1500 amino acids long while all the other organism in the tree are close to 5000 amino acids in length. BLOSUM62 most likely produced a tree where the lobster was the outgroup because its calculating the likelihood that a large amount of amino acids are completely missing in RYR in comparison to the other homologues.
References:
[1]phylogeny. (2014). In Encyclopædia Britannica. Retrieved from http://www.britannica.com/EBchecked/topic/458573/phylogeny
[2]Images of Phylogenetic trees from Understanding Evolution. (2011). University of California Museum of Paleontology. Retrieved from
http://bcrc.bio.umass.edu/intro/manual/index.php/Studying_Evolutionary_Relationships
[3]Fassler J, Cooper P. BLAST Glossary. (2011 Jul 14). In: BLAST® Help [Internet]. Bethesda (MD): National Center for
Biotechnology Information (US). Retrieved from http://www.ncbi.nlm.nih.gov/books/NBK62051/
[1]phylogeny. (2014). In Encyclopædia Britannica. Retrieved from http://www.britannica.com/EBchecked/topic/458573/phylogeny
[2]Images of Phylogenetic trees from Understanding Evolution. (2011). University of California Museum of Paleontology. Retrieved from
http://bcrc.bio.umass.edu/intro/manual/index.php/Studying_Evolutionary_Relationships
[3]Fassler J, Cooper P. BLAST Glossary. (2011 Jul 14). In: BLAST® Help [Internet]. Bethesda (MD): National Center for
Biotechnology Information (US). Retrieved from http://www.ncbi.nlm.nih.gov/books/NBK62051/
***Cover photo accessed from http://scientopia.org/blogs/guestblog/tag/phylogeny/
Site created by: Mercede Davis
Email contact: [email protected]
Date last updated: 5/14/14
Genetics 564, University of Wisconsin-Madison