This website was created as an assignment for Genetics 564 at the University of Wisconsin-Madison
Homology of Proteins
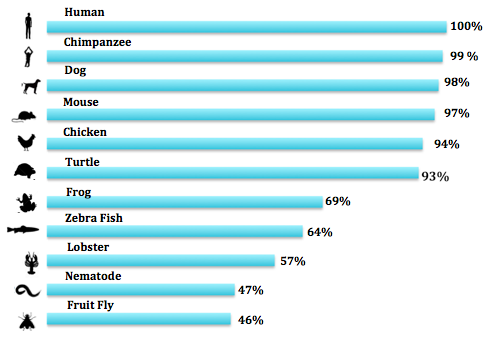
The homology of proteins is extremely important for the study of human diseases. Homologous proteins are very likely to function in the same manner because of their similar amino acid sequence and size. Being able to identify a protein homologue is necessary for the use of model organisms. In recent years, politicians have criticized scientists for spending "unnecessary" amounts of money studying model organisms such as fruit flies. Little do they know that something as simple as a fruit fly shares an enormous amount of its DNA with humans and other organisms. The homology between a model organism such as a fruit fly and a human make it possible to research human genetic disorders both ethically, financially and in a timely manner.
Homology of Ryanodine Receptor 2
I decided to keep the same organisms that were already identified as gene homologues and ran them through a protein BLAST. Just like the mRNA BLAST used for the RYR2 gene, protein BLAST searches for alignments in amino acid sequence and provides a percent identity and an E value in its output. I also calculated a percent identity using both a MUSCLE (Multiple Sequence Alignment Comparison by Log Expectation) and Clustal Omega protein alignment. Both Clustal Omega and MUSCLE use different algorithms for completing their alignments. Clustal Omega is the newest alignment tool in the Clustal Family and is said to be far superior and accurate when making larger alignments in comparison to other programs[1].
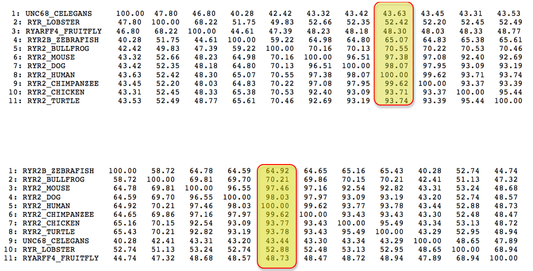
Clustal Omega and MUSCLE Alignment Matrices
|
|
Both Clustal Omega and MUSCLE are able to calculate percent identity based off of their different alignments. Each program creates a matrix of values by comparing each amino acid sequence to each other and determining its percent identity[1]. Highlighted in yellow are the columns that have calculated the percent identity of humans and each individual animal. As you can see, the calculated percent identities from both Clustal Omega and MUSCLE are very similar. In comparison to BLAST, Clustal Omega and MUSCLE's percent identity were more precise, calculating the percent identity to the hundredth percent. Also, BLAST calculated a percent identity of C. elegans and Lobster much higher than the other two programs. To view the actual amino acid alignments, click the buttons on the left.
|
Analysis
By analyzing the results of the three different programs, it is obvious the RYR2 protein is highly conserved in mammals. Chimpanzees, dogs, and mice all have extremely high percent identities with the human RYR2 protein. Even reptiles and birds, represented by the chicken and the turtle, have conserved an enormous amount of the amino acid sequence. As we get further away and out of the vertebrate category, this number decrease, but still presents evidence that RYR2 has been very well preserved in many different phyla and throughout the Animalia Kingdom.
Homologue References

Chimpanzee (Pan troglodytes) Ryanodine Receptor 2
Accession number:XP_514296 GI Number:410034672 FASTA Percent Identity:99% E Value: 0.0 |

Dog (Canis lupus) Ryanodine Receptor 2 isoform X2
Accession Number: XP_005618837 GI Number:545494606 FASTA Percent Identity: 98% E Value: 0.0 |

Turtle (Chrysemys picta) Ryanodine Receptor 2
Accession Number: XP_005291316 GI Number:530595513 FASTA Percent Identity: 93% E Value: 0.0 |

Zebra Fish (Danio rerio) Ryanodine Receptor 2b
Accession Number: XP_00192113 GI Number:189530774 FASTA Percent Identity: 64% E Value: 0.0 |

Lobster (Homarus Americanus) RYR
Accession Number: AAC06013 GI Number:2970625 FASTA Percent Identity: 57% E Value: 0.0 |
Nematode (Caenorhabditis elegans) UNC-68
Accession Number:NP_00125607 GI Number: 392919361 FASTA Percent Identity: 47% E Value: 0.0 |

Fruit Fly (Drosophila melanogaster) Ryanodine Receptor 44F Isoform A
Accession Number:NP_476991 GI Number: 17352465 FASTA Percent Identity: 46% E Value: 0.0 |
References:
[1]Jurate Daugelaite, Aisling O' Driscoll, and Roy D. Sleator, “An Overview of Multiple Sequence Alignments and Cloud
Computing in Bioinformatics,” ISRN Biomathematics, vol. 2013, Article ID 615630, 14 pages, 2013.
doi:10.1155/2013/61563. www.hindawi.com/isrn/biomathematics/2013/615630/
IMAGES:
http://lovendar.com/articles/Top-10-Common-Sings-of-a-Happy-Couple
http://uncyclopedia.wikia.com/wiki/File:Chimpanzee-picture.jpg
http://advocacy.britannica.com/blog/advocacy/2011/09/beagles-deserve-better/
http://wandervogeldiary.wordpress.com/2012/07/29/
http://en.wikipedia.org/wiki/Painted_turtle
http://todayistheday-bio156.blogspot.com/2013/04/chicken-legs.html
http://www.bionalogy.com/role_of_genetic_information.htm
http://www.lobsterhelp.com/types-of-lobster.html
http://www.theradzoo.com/meet-the-animals/frogs-toads/african-bullfrog/
http://www.elvesys.com/fast-temperature-controller-for-microfluidic-chip
http://www.domyownpestcontrol.com/fruit-flies-c-149.html
Site created by: Mercede Davis
Email contact: [email protected]
Date last updated: 5/14/14
Genetics 564, University of Wisconsin-Madison
Email contact: [email protected]
Date last updated: 5/14/14
Genetics 564, University of Wisconsin-Madison