This website was created as an assignment for Genetics 564 at the University of Wisconsin-Madison
Protein Interaction Networks
Protein interactions are characterized by the ability of a protein to bind to each other to perform a designated function. These interactions are essential in every function of an organism and can be used to understand what is happening in an organism at a molecular level. Protein-protein interactions can often be something hard to identify due to their ever-changing expression levels that are affected by many different environmental factors or other biological process occurring within an organism. Luckily, scientists have developed methods that are are able to determine binding partners of a protein such as performing a Yeast two hybrid, TAP tagging, or coimmunoprecepitation. Binding partners can often provide scientists information about the function of their protein of interest and can often lead to new avenues for further research. Results from these experiments can be visualized as an interacting web of proteins. All webs created for RYR2 protein interactions below were created using STRING 9.1 [1].
Which Proteins Interact with RYR2?
STRING 9.1 was used to determine interacting partners for RYR2. STRING 9.1 creates protein interaction networks based on genomic context, high-throughput experiments, conserved coexpression, and previous knowledge from literature[1].
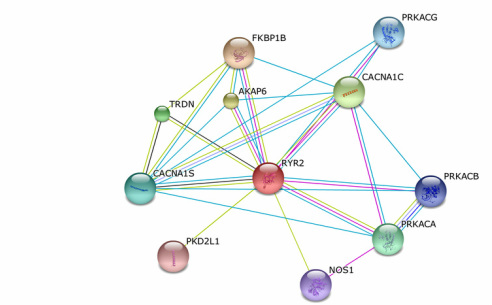
To the left is a protein interaction network for human RYR2 created by STRING 9.1. As you can see, RYR2 interacts with many different proteins. By understanding what the interacting proteins do, we can better understand how RYR2 is functioning and what processes it is involved in.
To understand what processes the interacting proteins of RYR2 are involved in, I filtered the protein network by the three gene ontology categories; cell component, molecular function, and biological processes.
To understand what processes the interacting proteins of RYR2 are involved in, I filtered the protein network by the three gene ontology categories; cell component, molecular function, and biological processes.
|
Cell Component
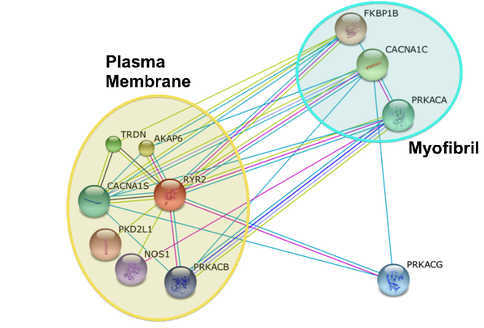
As seen on the Gene Ontology page, RYR2 is involved in many different cellular components. RYR2's binding partners are involved in many different cellular components as well. Seen to the right is RYR2's String protein interaction network. I have highlighted two groups of proteins, ones localizing to plasma membranes and the other localized to myofibrils, or muscle fibers. It is easy to see that the three proteins localized in the myofibril have an enormous amount of interactions with the membrane proteins suggesting collectively, the plasma membrane proteins are affecting muscle tissue in the heart [2]. |
|
Biological processes
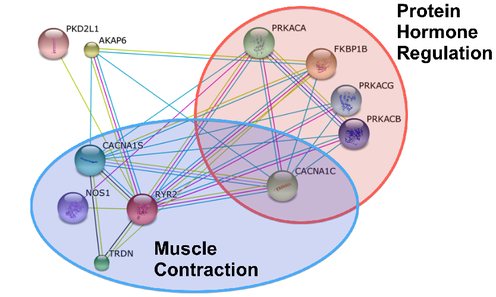
In Figure 2, the String is reorganized by biological processes. Two processes I highlighted were protein hormone regulation and muscle contraction. Interestingly enough, the three proteins located in the myofibrils in Figure 1 are in the protein hormone regulation category. Other interesting biological processes included insulin regulation, lipase regulation, and response to epinephrine [2]. |
|
Molecular function
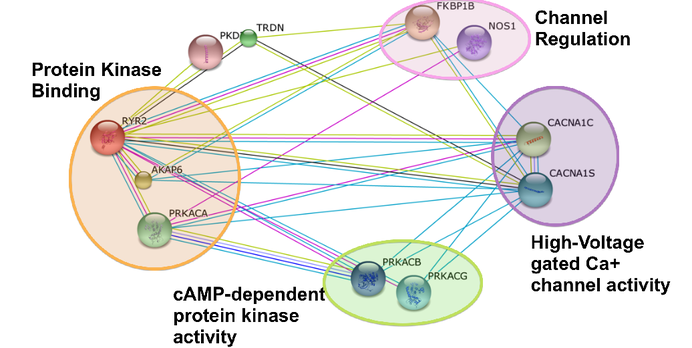
Figure 3 highlights a few of the molecular functions of RYR2's binding partners. Many of the molecular functions have to do with regulation of the calcium ion channel and its activity or calcium binding[2]. |
Analysis
As you can see, STRING was able to identify many different proteins that interact with RYR2. Many of the proteins identified had functions that aid in RYR2's overall function. Both CACNA1S and CACNA1C are involved in voltage gated calcium ion processes such as muscle contraction, but are also involved in horomone and neurotransmitter release and gene expression which I thought was very interesting. Both PRKACB and PRKACG are responsible for phosphorylation of many different substrates in the cytoplasm[2]. This makes sense because RYR2 requires phosphorylation to function[3]. AKAP6 is responsible for anchoring protein kinases into the sacroplasmic reticulum and is also thought to be involved in the assembly of the of the calcium ion complex. This could possibly suggest that AKAP6 is involved in RYR2's localization to the sacroplasmic membrane. One protein interaction that I thought was interesting was RYR2 and PKD2L1, a protein that causes Polycystic Kidney Disease. Though PKD2L1 is thought to be a transducer of calcium mediated signaling, it is interesting that PKD2L1 does not interact with any of the other proteins in the interaction network. I also find it interesting that so many proteins were involved in protein, gene, and neurotransmitter regulation which goes to show that RYR2's function is dependent on many different proteins and that calcium regulation is complicated process [2].
References:
[1]STRING-Known and predicted protein-protein interactions. Accessed April 7, 2014 from http://string-db.org/
[2] RYR2 protein STRING interaction web. Accessed April 12, 2014 from http://string-db.org/newstring_cgi/show_network_section.pl
[3] Adenylyl Cyclase. Retrieved March 7, 2013, from http://webspace.ship.edu/cgboer/genpsyneurotransmitters.html
[1]STRING-Known and predicted protein-protein interactions. Accessed April 7, 2014 from http://string-db.org/
[2] RYR2 protein STRING interaction web. Accessed April 12, 2014 from http://string-db.org/newstring_cgi/show_network_section.pl
[3] Adenylyl Cyclase. Retrieved March 7, 2013, from http://webspace.ship.edu/cgboer/genpsyneurotransmitters.html
Site created by: Mercede Davis
Email contact: [email protected]
Date last updated: 5/14/14
Genetics 564, University of Wisconsin-Madison
Email contact: [email protected]
Date last updated: 5/14/14
Genetics 564, University of Wisconsin-Madison