This website was created as an assignment for Genetics 564 at the University of Wisconsin-Madison
What is Homology?
Homology is the similarity between organisms based on their decent from a common ancestor. These similarities can be at the structural level, displaying the commonalities of skeletal and other morphological features between species. Homology can also be applied at the molecular level, where scientists can compare both genetic code and protein amino acid sequence to determine how specific genes have been conserved or have changed throughout evolution [1]. The study of both gene and protein homology is very important for the study of human diseases. Scientists and doctors can use these similarities between human genes and proteins and those of model organisms to research and better understand human genetics and it's relation to disease. They are able to use organisms with a high percent identity as model organisms for research to give better insight to how the disease functions within a human.
Homology of the RYR2 Gene
To determine the homology of the RYR2 gene seen below, a BLAST (Basic Local Alignment Search Tool) was run. BLAST uses an alignment algorithm that takes the desired nucleotide or amino acid sequence of an organism and compares it to the reference genomes of other organisms that are stored within its database. Using these alignments, BLAST is able to search for specific gene or amino acid sequence within the organism you are investigating and determine the percent identity in respect to your original gene or protein. To find RYR2 homologues, the nucleotide sequence of the human RYR2 mRNA (only the protein coding regions of the DNA) was used to search for other organisms with similar RYR2 genetic code. I decided to use the mRNA sequence instead of the DNA sequence because the DNA nucleotide sequence of the human RYR2 gene is extremely long and it takes a substantial amount of time for BLAST to make alignments with the introns (non-coding regions) still in the sequence. Also, by using the mRNA sequence, all of the introns have already been spliced out of the sequence which leaves us with only region that codes the Ryanodine Receptor 2 protein. BLAST provides an E Value with each homologue in which the smaller the E Value, the more statistically significant the results are [2].
|
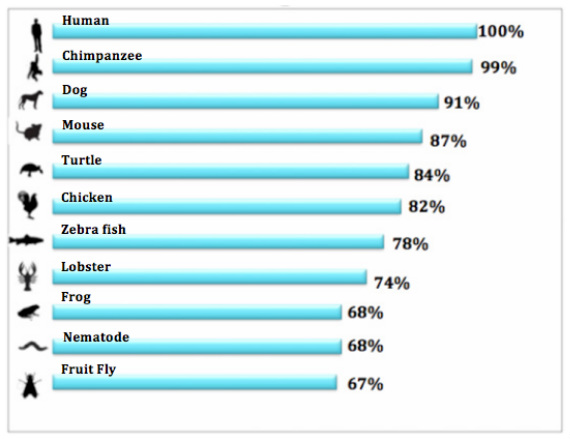
Figure 1: This figure was created to demonstrate visually the percent identity between mRNA sequences of different organisms and the human RYR2 mRNA sequence. These percent identities were calculated by BLAST. More detail on each homologue can be found below in the Homologue References section. |
Analysis
When comparing the human RYR2 gene to other organisms from BLAST, you can easily see that the RYR2 gene is highly conserved in mammals. The highest percent identity was seen in chimpanzees, which is reasonable because the chimpanzee is known for being genetically very similar to the human. Dog and mouse were the next highest percent identities in the organisms I BLASTed. This is great, because both dog and mouse have been used as a model organism for studying CPVT [4]. As we diverge from mammals, the percent identities decreases, but still maintains high conservation. RYR2 is only expressed in cardiac tissue but is closely related to the Ryr1 and Ryr3 genes that code Ryanodine Receptor proteins in non-cardiac tissues such as skeletal muscle. This is why BLAST found homologues in organisms without a traditional heart, such as C. Elegans. Unc-68 in C. Elegans acts in muscle tissue and controls muscle contraction involved in locomotion. Even though Unc-68 doesn't function in the same tissue and thus doesn't have the exact same function on a macro level, Unc-68 is still biochemically comparable to RYR2 and is homologous to all three of the genes found in the Ryanodine Receptor family[3]. BLAST was used against, plants, fungi, bacteria, and archea, but no homologues were found, suggesting that maybe the RYR2 gene developed after animals diverged from these other organisms.
Homologue References

Chimpanzee (Pan troglodytes) RYR2
Accession Number: XM_514296 GI Number: 410034671 FASTA Percent Identical: 99% E Value: 0.0 
Mouse (Mus musculus) RYR2
Accession Number: NM_023868 GI Number:124430577 FASTA Percent Identical: 87% E Value: 0.0 
Chicken (Gallus gallus) RYR2
Accession Number: XM_419553 GI Number:513175707 FASTA Percent Identical:82% E Value: 0.0 
Lobster (Homarus Americanus) RYR
Accession Number: AF051936 GI Number: GI:2970624 FASTA Percent Identical:74% E Value: 9e-82 
Nematode (Caenorhabditis elegans) Unc-68
Accession number: NM_001269146 GI Number: 453232387 FASTA Percent Identical: 68% E Value: 1e-45 |

Dog (Canis lupis) RYR2
Accession Number: XM_005618780 Gi Number:545494605 FASTA Percent Identical: 91% E Value:0.0 
Painted Turtle(Chrysemys picta) RYR2
Accession Number: XM_005291259 GI Number: 530595512 FASTA Percent Identical: 84% E Value: 0.0 
Zebrafish (Danio rerio) Ryr2a
Accession Number: XM_002667555 GI Number: 528496857 FASTA Percent Identical: 78% E Value: 0.0 
Bull Frog (Rana catesbeiana) RYR3
Accession Number: D21071 GI Number:29501271 FASTA Percent Identical: 68% E Value: 0:0 
Fruit Fly (Drosophila melanogaster) Rya-r44F
Accession Number: NM_001259281 GI Number:386767558 FASTA Percent Identical: 67% E Value 1e-136 |
References:
[1]homology. (2014). In Encyclopædia Britannica. Retrieved from
http://www.britannica.com/EBchecked/topic/270557/homology
[2]Gene Reviews. (2013). Electronic references. Retrieved January 31, 2014, from
http://www.ncbi.nlm.nih.gov/books/NBK1289/
[3]Unc-68 (gene). WormBase:Nematode Information Resource. Retrieved from http://www.wormbase.org/species/c_elegans/gene/WBGene00006801
[4]OMIM. (2011). Electronic References. Retrieved January 28, 2014 from http://www.omim.org/entry/180902
IMAGES:
http://lovendar.com/articles/Top-10-Common-Sings-of-a-Happy-Couple
http://uncyclopedia.wikia.com/wiki/File:Chimpanzee-picture.jpg
http://advocacy.britannica.com/blog/advocacy/2011/09/beagles-deserve-better/
http://wandervogeldiary.wordpress.com/2012/07/29/
http://en.wikipedia.org/wiki/Painted_turtle
http://todayistheday-bio156.blogspot.com/2013/04/chicken-legs.html
http://www.bionalogy.com/role_of_genetic_information.htm
http://www.lobsterhelp.com/types-of-lobster.html
http://www.theradzoo.com/meet-the-animals/frogs-toads/african-bullfrog/
http://www.elvesys.com/fast-temperature-controller-for-microfluidic-chip
http://www.domyownpestcontrol.com/fruit-flies-c-149.html
[1]homology. (2014). In Encyclopædia Britannica. Retrieved from
http://www.britannica.com/EBchecked/topic/270557/homology
[2]Gene Reviews. (2013). Electronic references. Retrieved January 31, 2014, from
http://www.ncbi.nlm.nih.gov/books/NBK1289/
[3]Unc-68 (gene). WormBase:Nematode Information Resource. Retrieved from http://www.wormbase.org/species/c_elegans/gene/WBGene00006801
[4]OMIM. (2011). Electronic References. Retrieved January 28, 2014 from http://www.omim.org/entry/180902
IMAGES:
http://lovendar.com/articles/Top-10-Common-Sings-of-a-Happy-Couple
http://uncyclopedia.wikia.com/wiki/File:Chimpanzee-picture.jpg
http://advocacy.britannica.com/blog/advocacy/2011/09/beagles-deserve-better/
http://wandervogeldiary.wordpress.com/2012/07/29/
http://en.wikipedia.org/wiki/Painted_turtle
http://todayistheday-bio156.blogspot.com/2013/04/chicken-legs.html
http://www.bionalogy.com/role_of_genetic_information.htm
http://www.lobsterhelp.com/types-of-lobster.html
http://www.theradzoo.com/meet-the-animals/frogs-toads/african-bullfrog/
http://www.elvesys.com/fast-temperature-controller-for-microfluidic-chip
http://www.domyownpestcontrol.com/fruit-flies-c-149.html
Site created by: Mercede Davis
Email contact: [email protected]
Date last updated: 5/14/14
Genetics 564, University of Wisconsin-Madison