This website was created as an assignment for Genetics 564 at the University of Wisconsin-Madison
Specific Aims and Conclusions
Catecholaminergic Polymorphic Ventricular Tachycardia is an extremely deadly genetic disorder. Over 30 percent of people with CPVT die before the age of thirty from cardiac arrest, many of which are completely unaware that they even had a heart condition [1,2]. Though once diagnosed individuals with CPVT can be treated, it requires major life changes and many times surgery is necessary as well [2]. The primary goal of these specific aims is to better understand how RYR2 mutants function differently within different regions of the heart. Since CPVT is only a ventricular arrhythmia disorder [1,2], it is logical to conclude that RYR2 functions differently between the atria and the ventricles. Identifying these key differences in RYR2 function could help scientists better understand RYR2's role in the disorder and aid in developing better techniques for treatment or help find a cure.
Aim #1
First, I wanted to compare human RYR2 to mouse RYR2 to determine the similarities between the two genes. This is an extremely important step when using model organisms for research. I predicted that RYR2 would be extremely conserved in the mouse model since RYR2 is necessary for heart function. Since it is an important gene for such a vital organ, this is a very logical hypothesis. To investigate this aim, I used BLAST to determine percent identity between the two RYR2 proteins.
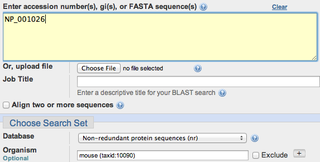 Human RYR2 BLASTed against mouse [3]
Human RYR2 BLASTed against mouse [3]
BLAST determined that the RYR2 protein of the mouse was 97% identical to the human RYR2 protein[3]. This is not surprising since mice and humans share the same type of circulatory system since they are both mammals. BLAST also showed that the mouse RYR2 protein is only one amino acid less than the human amino acid[3]. Not only is RYR2 highly conserved in our mouse model, but it is also essentially the same length. This suggests that RYR2 is going to function in the same manner as it would in the human making it a great model to work with for further research. CLUSTAL OMEGA was also used and determined a 97.46% identify for mouse RYR2 and human RYR2. It also showed that there are very few amino acid differences between the two protein sequences [4].
Aim #2
Next, I wanted to investigate how RYR2 is expressed differently between atria and ventricles. To do this I would perform a microarray on both wild-type and RYR2 mutant mice. A microarray would be conducted both before and after exercise to better understand how RYR2 functions in CPVT individuals in comparison to normal individuals. Using GEO, I found an RYR2 profile showing expression differences of RYR2 in the atria and the ventricles of the common rat, a close relative of the mouse.
Next, I wanted to investigate how RYR2 is expressed differently between atria and ventricles. To do this I would perform a microarray on both wild-type and RYR2 mutant mice. A microarray would be conducted both before and after exercise to better understand how RYR2 functions in CPVT individuals in comparison to normal individuals. Using GEO, I found an RYR2 profile showing expression differences of RYR2 in the atria and the ventricles of the common rat, a close relative of the mouse.
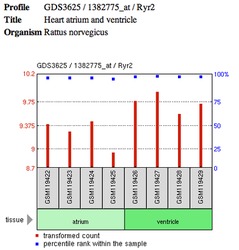 RYR2 Profile from GEO [5] RYR2 Profile from GEO [5]
From this profile, there is a clear discrepancy in expression levels between the two regions. I would expect to see this same pattern in our mouse model. This profile of the rat leads the to believe that this expression level difference is a contributing factor to the ventricular aspect CPVT [5]. I would expect that these expression levels increase during exercise for the wild type mouse, but only the atria RYR2 expression level will increase in the RYR2 mutant mouse after exercise. My logic behind this prediction is that RYR2 expression in the ventricles of the mutant mouse is not able to keep up with the demand of an increasing heart rate and essentially starts to malfunction. If this hypothesis is found to be correct, it would explain why the phenotype is only seen in the ventricles during increased heart rate.
Aim #3 Due to the the expression level differences of RYR2 I expect to see in the microarray performed in Aim #2, I believe that there are other proteins involved in RYR2's function. These proteins interacting with RYR2 are either inhibiting RYR2 mutations from causing an arrhythmia in the atria or are promoting the arrhythmia phenotype of the ventricles of the heart. To further investigate this, I would add a TAP-tag to RYR2 in wild type mice and in RYR2 mutant mice. 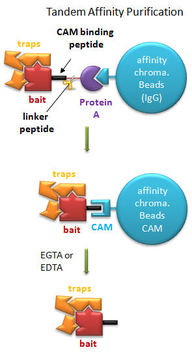 Protein complex isolation via TAP-tag Protein complex isolation via TAP-tag
The TAP-tag would be used to pull down RYR2 and other proteins involved in the calcium-ion channel complex [6]. The isolated complex would then be analyzed via mass spectrometry to determine which proteins are directly interacting with RYR2 and at what quantities they are being expressed. Samples would be taken from both atria and ventricles before and after exercise to understand how RYR2 is functioning in CPVT producing conditions. My hypothesis is that when analyzed via mass spectrometry, either there are interacting partners present in the atria samples that are not present in the ventricular sample or vice versa or some proteins are expressed at extremely different proportions between the two regions. Proteins identified with this peculiar characteristic potentially could be altering RYR2 function between the atria and ventricles. I also predict that there will be a difference between pre-exersize samples and post-exercize samples when comparing wild type and RYR2 mutant mice. These differences could also help us identify how RYR2 mutants are not functioning properly under this condition and what proteins are preventing this dysfunction in the atria.
|
Conclusions
Through these aims we will hopefully be able to better understand how RYR2 is functioning differently throughout the heart and how mutations in RYR2 affects that function. Proteins involved in altering RYR2's function and expression level will be identified and this identification could lead to other avenues for comprehensive CPVT research and understanding of the disorder. By determining the cause of RYR2's altered function, we will be able to better understand mechanistically how CPVT develops and this knowledge could provide insight for developing other techniques for treatments or aid in finding a cure.
Future Research
After performing the experiments outlined in the specific aims, I would want to further investigate proteins involved in RYR2's function. I think it would be interesting to return the the microarray performed in Aim #2 and look for expression level changes of the proteins identified in Aim #3 and determine how those expression levels are changing in conjunction with RYR2. Also, since CPVT is caused by over 70 point mutations, it would be interesting to further classify each mutation and determine if a CPVT causing mutation in RYR2 is disrupting its interaction with the proteins identified with Aim #3.
| cpvt_final_talk.pptx | |
| File Size: | 4505 kb |
| File Type: | pptx |
References:
[1]Paivi J. Laitinen, Kevin M. Brown, Kirsi Piippo, Heikki Swan. (2001). Mutations in Cardiac Ryanodine Receptor (RyR2) Gene
in Familial Polymorphic Ventricular Tachycardia. Circulation. 103. 485-490. doi: 10.1161/01.CIR.103.4.485
http://circ.ahajournals.org/content/103/4/485.full.pdf+html
[2]SADS Foundation. (2008). Electronic references. Retrieved February 1, 2014 from
http://www.sads.org/library/cpvt#.UuQigDl6i1s
[3]Basic Local Alignment Search Tool. NCBI. Accessed May 4, 2014 from http://blast.ncbi.nlm.nih.gov/Blast.cgi
[4]Clustal Omega. EBI. Accessed May 4, 2014 from http://www.ebi.ac.uk/Tools/msa/clustalo/
[5]RYR2 GEO profile. Accessed April 23, 2014 from http://www.ncbi.nlm.nih.gov/geo/tools/profileGraph.cgi?ID=GDS3625:1382775_at
[6]Introduction to Tandem Affinity Purification. (2012). Bitesize Bio. Accessed May 7, 2014 from http://bitesizebio.com/8208/an-introduction-to-tandem-affinity-purification/TAP-tag Image- http://employees.csbsju.edu/hjakubowski/classes/ch331/bind/TandemAffPurif.jpg
Site created by: Mercede Davis
Email contact: [email protected]
Date last updated: 5/14/14
Genetics 564, University of Wisconsin-Madison
Email contact: [email protected]
Date last updated: 5/14/14
Genetics 564, University of Wisconsin-Madison
